Molecular modeling of ion channels
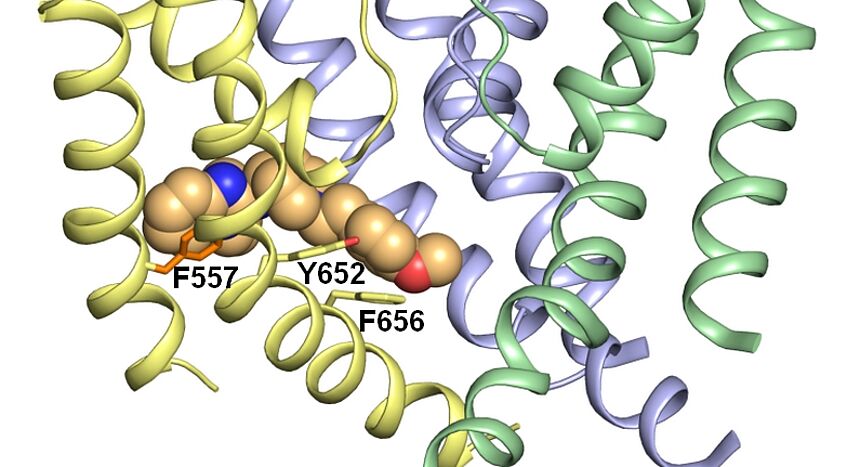
(Foto: Anna Weinzinger)
We use computational methods (including homology modelling, molecular dynamics simulations, docking investigations etc.) to explore the relationship between structure, function and dynamics of ion channels. We currently focus on several distinct ion channel families including K+, Ca2+ and Nav channels. The recent breakthrough in x-ray crystallography and cryo-electron-microscopy, enables detailed insights into the three-dimensional architecture of more and more of these channels, allowing detailed insights into their structure. Although, there is an increasing amount of structural information available, many questions, particular concerning the dynamics associated with gating, selectivity, conductance and ligand binding remain unanswered. Computer simulations allow us to simulate the motions of these proteins and to explore the relationship between (static) structure and dynamic function.
Our main research questions are (i) to understand how ion channels are gated (ii) how they are regulated by physiological ligands, (iii) how inherited gain-or loss-of-function mutations affect normal function, and (iv) how small molecule modulation affects channel action (including off-target effects). Our multi-disciplinary ranges from state-of-the-art in silico methods such as molecular dynamics simulations, enhanced sampling techniques, free energy calculations and computational electrophysiology and in close collaboration with experimental labs we perform (bio)chemical, pharmacological and biophysical studies, which enable us to unravel the inner workings of ion channels.

(Foto: Michael Bründl)
Selected publications:
A complete list of publicatons can be found here
Chen, X, Garon, A, Wieder, M, Houtman, MJC, Zangerl-Plessl, EM, Langer, T, van der Heyden, MAG, Stary-Weinzinger, A. (2019) Computational identification of novel Kir6 channel inhibitors. Front. Pharmacol. in press https://doi: 10.3389/fphar.2019.00549
Dierich, M, van Ham, W.B., Stary-Weinzinger, A, Leitner M.G. (2019) Histidine at determines the low quinine sensitivity of ether-à-go-go (Eag) channel superfamily member Kv12.1. Br J Pharmacol. in press https://doi: 10.1111/bph.14693
Houtman, M, Chen, X, Quile, M, Duran, K, van Haaften, G, Stary-Weinzinger, A, van der Heyden, M.A.G. (2019) Glibenclamide and HMR1098 normalize Cantú Syndrome-associated gain-of-function currents. J Cell and Mol Med. in press
Chen, X, Bründl, M, Friesacher, T, Stary-Weinzinger, A. (2020) Computational Insights Into Voltage Dependence of Polyamine Block in a Strong Inwardly Rectifying K + Channel. Front. Pharmacol. in press www.frontiersin.org/articles/10.3389/fphar.2020.00721/full
Zangerl-Plessl, EM, Lee, SJ, Maksaev, G, Bernsteiner, H, Ren, F, Yuan, P, Stary-Weinzinger, A, Nichols, CG. (2020) Atomistic basis of opening and conduction in mammalian inward rectifier potassium (Kir2.2) channels. J Gen Physiol. in press rupress.org/jgp/article/152/1/e201912422/132552/Atomistic-basis-of-opening-and-conduction-in
Preprints:
Bründl, M, Pellikan, S, Stary-Weinzinger, A. (2020) Simulating PIP2-induced gating transitions in Kir6.2 channels. www.biorxiv.org/content/10.1101/2021.05.20.445012v1
Current Funding:
FWF doctoral program "Molecular Drug Targets" grant W1232, 2010 - 2023
e-rare2: "Sulfonylurea to treat Cantú syndrome", 2015 - 2019, FWF
ÖAW „DOC Fellowship Programme“, 2021-2023, Austrian Academy of Sciences
Former group members:
Xingyu Chen (MolTag PhD student)
Eva Hellsberg (MolTag PhD student, Ecker group, 1year internship)
Milica Gacesa (master student)
Niko Schramböck (master student)
Nabil Mohamed (master student)
Simone Strohmaier (bachelor student)
Sarala Pellikan (PhD student)
Peter Gröschl (master student)
Emina Selimovic (master student)
Wironia Gadalla (master student)
Renate Hofer (master student)
Sara Eskander (master student)
Melina Tindl (master student)
Lisa Pammer (master student)
